Clinical Diagnosis for Computer Scientists: Machines reading patient records

Experienced doctors are better doctors. We’d all fight over that doctor that has seen a hundred cases like yours last week. Alas, most doctors haven’t. Thankfully they are routinely recording their experiences in electronic health records. The dream of EHR research is to create insights from millions of patient records, something humans can’t really do.
How far can this go?
Earlier this year the Tatonetti Lab conducted the largest heritability study ever. They estimated the heritability of 500 traits with very high confidence. Their dataset, or even the underlying method, could be used to identify patients at genetic risk of disease. Their code is on github.
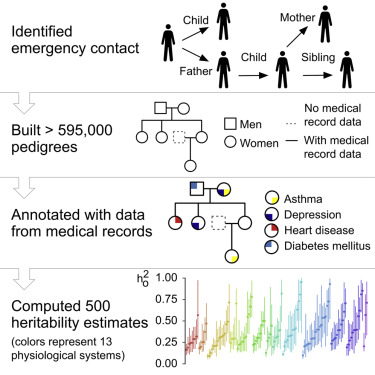
 Their study is special because it’s a genius hack! Until now geneticists have been gathering thousands of people and conducting limited studies to find out how a few traits are inherited. Instead, this team used emergency contact information to figure out how 3.5 million patients are related - quite possibly a GDPR nightmare if conducted in the EU. The data was already there! Then they use an old genetics trick called linkage analysis, to model how the traits pass through the family trees. I recommend the original SOLAR paper if you’re craving some stats.
Their study is special because it’s a genius hack! Until now geneticists have been gathering thousands of people and conducting limited studies to find out how a few traits are inherited. Instead, this team used emergency contact information to figure out how 3.5 million patients are related - quite possibly a GDPR nightmare if conducted in the EU. The data was already there! Then they use an old genetics trick called linkage analysis, to model how the traits pass through the family trees. I recommend the original SOLAR paper if you’re craving some stats.
Google took a more general angle, proposing a deep architecture to predict anything you want from medical records. This single architecture could predict (quite well) a patient’s mortality, readmission, length of stay and even give some insight into the discharge diagnosis. What’s impressive here is that their engineers didn’t need any medical/biological knowledge, or to even clean the 200k medical records. It just worked. And better than ever before.
But what can it not do
There is one area that is stumping everyone: diagnosis prediction. Figuring out which disease a patient has from a doctor’s findings is something even doctors struggle to do. To my surprise, Netflix and the NYT have just launched a series on this very problem. Even Google’s deep architecture fell short on this task, getting an F1 score of 0.4-0.41. This is In fact, not much more compelling than the 0.4 precision score DeepPatient achieved two years ago.
Why are we so bad at this?
Firstly, diagnosis prediction is complex. Let’s consider just one knowledge base that has been used for diagnosis tools, HPO. It contains around 150,000 relationships between clinical findings and disease. Each disease has on average 14 clinical features, each of which:
- can have up to 30 synonyms
- may or may not appear in the patient, see penetrance
- may be more likely to occur at a particular age
And this is one small, structured database. Imagine the complexities that lie in the patient records! And we’re trying to predict one or two diseases out of about 8000.
This wouldn’t be a problem for deep learning if it wasn’t for the fact that a large portion of these diseases are rare. How are you going to get the data to train a network when fewer than 1 in 2,000 of the population are affected? You will quickly run into the sparse data problem.
Cramming knowledge
For this task experience is not enough, we need to cram like a 3rd year medic. And that’s an exciting challenge. Experts spend years understanding the biochemical mechanisms of disease to eventually figure out what’s going on with a patient. I believe machines will do the same in a few hours. And this is a new paradigm of machine learning. Models would not only be learning from experience. They would be applying knowledge too.
As far as I know, no one has done this for diagnosis prediction (IBM doesn’t count). But it has sprung up in research under the guise of Distant Supervision and Data Programming. I suspect methods like this will push EHR over this next big hurdle.
This is the first in a series on Clinical Diagnosis for Computer Scientists. If this goes well, I am up for writing posts on Genetic Diagnosis, Zero-Knowledge Diagnosis (patient data sharing) and Personalised Clinical Trials.
If you liked this article maybe you will also find interesting the following in-depth articles about other rare diseases, like A Day In The Life Of...Dr Pradeep Ravichandran 10 million patient-disease scans: 5 lessons we’ve learned Regulation - Far More Than A Tick-Box Retrospective analysis suggests MendelScan saves 3.5 years of medical and financial burden in Behçets disease Mendelian among the most innovative companies of the year by "El Pais". Orphan Drugs – Meeting Global Unmet Health Needs